Did you see the news? A Christmas gift to Science.
DeepMind AI’s new version of a program called AlphaFold is creating waves in the scientific community. The program, backed by a truck load of computing power, can very accurately predict the exact folding and shape of proteins from their underlying amino acid sequences.
DeepMind, if you remember, is the company behind the AI program AlphaGo that made headlines back in 2016 when it beat Go’s reigning champion, 18 time European winner Mr Lee Sedol. Go, a game originating in China, is said to be one of the hardest strategy games in the world, requiring several different levels of logical thinking to succeed.
This time is different. AlphaGo was a test run, used to master a complex game. It showed the potential of AI software but lacked any significant real world application. Their most recent software, AlphaFold, has incredible potential in many scientific fields and leads the way for further innovation for AI applications in scientific research.
In this post, I'm going to try and explain why this is such an important breakthrough
I’ll go back to the basics for those who don’t have the scientific background and hopefully explain why this is such a relevant development in the field
Back to Basics - What are proteins?
Proteins are the building blocks of all cells and tissues in the human body. While some types of proteins are just purely for structure, most form important ‘biological machinery’. Proteins themselves are composed of chains of 20 different amino acids in different sequences. The different amino acids have different properties and are fully responsible for the final 3D structure of the protein.
Certain sequences of amino acids can form different types of what are called secondary structures, which are basically little structures inside the protein. Some examples of these are the alpha helix and the beta pleated sheet shown in their simple forms below:

Alpha Helix
Beta Pleated Sheet
For more detailed information on protein structure, click here.
These structures may occur multiple times during the protein structure, so these structures themselves will also have a certain position in the 3D space.
For example, here's the Haemoglobin protein
Haemoglobin is the protein in red blood cells that is able to capture oxygen and transport it around the body in the blood stream. The structure below is interactive – go ahead and rotate it around!
If you look carefully you can see many alpha helices (mini coils) all in different positions not only in the protein chain, but also in 3D space.
The problem of 3D structure has been around for more than 50 years
In 1972, Christian Anfinsen hypothesized in his Nobel Prize Speech that a protein’s 3D structure could be determined solely from it’s ‘1D’ amino acid sequence. Since then, scientists have been trying to predict exactly that with varying degrees of success as measured by the CASP assessment, a protein folding prediction ‘competition’ that happens every 2 years. This year there was a clear winner: AlphaFold 2.0.
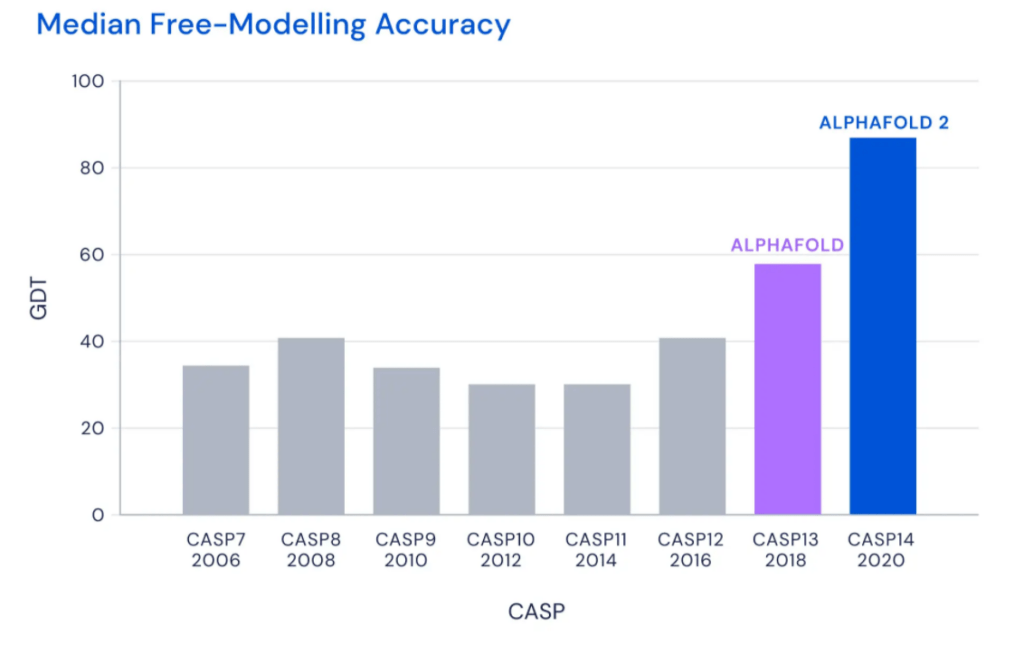
AlphaFold 2.0 leads previous winners in the dust with a median score of over 90 points. A score that high means the AlphaFold structure is incorrect by less than 1.6 Angstroms; the width of an atom.
Image taken from the following source: AlphaFold: a solution to a 50-year-old grand challenge in biology | DeepMind
Why is the 3D protein structure so important?
The exact shape or conformation of the protein is heavily involved in its function. In fact, most enzymes will not function at all even with a slight change to their 3D shape. A large number of these proteins in the body are enzymes or receptors; both of which are vastly important in bodily functions. Enzymes are the biological machinery responsible for reactions in metabolic pathways – pathways that break down nutrients to build up important molecules for life. Receptors are equally important for signaling in the body. In a previous post, I discussed the G Protein-Coupled Receptor in allergic reactions, take a look!
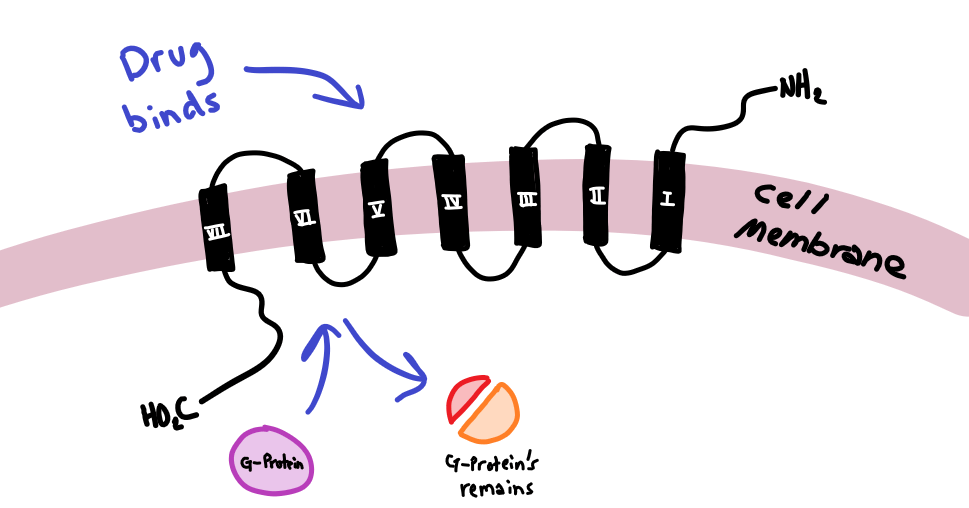
Here, a drug binds to a receptor on the outside of a cell and causes a signal to be transmitted on the inside of a cell using something called a G-Protein.
Why do we care?
Proteins and enzymes are the most commonly used drug targets. Almost all drugs on the market work via their interaction with an enzyme or receptor – two types of proteins. These drugs usually work because the structure of the drug molecule fits into a site on these enzymes and receptors perfectly like a hand in glove, so it can cause some effect how the protein does its job with the aim of treating a disease or symptom.
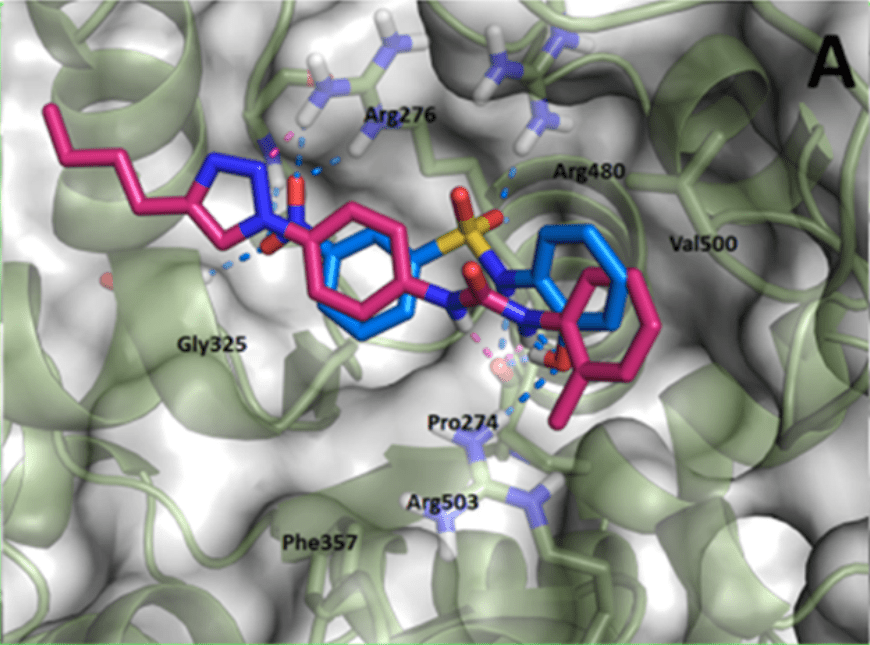
For example, on the left the binding of two (separate) potential drug molecules in the binding site of an enzyme is shown. In this particular paper, the 3D shape of the enzyme was actually used to design one of these drug molecules to fit perfectly in the enzyme.
Back to the AlphaFold software, what's the hype?
In order to make drugs to treat diseases, it is very important to know the physical shape and conformation of these proteins. Without it, scientists are left testing for drugs almost blindly, without knowing even where or how they work.
At the moment, many drugs are found through a technique called High Throughput Screening, where thousands of structurally similar molecules are tested for their biological activity on what is essentially a whim in the hopes of finding something with a biological activity that can be investigated.
Roughly one in 3,000 molecules will show some biological effect, but that doesn’t mean that the biological effect is useful or strong enough to be used! Then, a significant amount of drug candidates that make it through those criteria will show an unwanted side effect or toxicity which often means the end of the road for that particular drug candidate.
Knowing the 3D structures of proteins without painstaking work with expensive equipment has the potential to flip the process on it’s head – molecules could be accurately designed to have a structure that fits into the protein, speeding up the process of finding drug candidates for serious diseases.
How do we currently find 3D protein structures?
At the moment, protein structures are discovered through X-Ray crystallography, Nuclear Magnetic Resonance or cryo-electron microscopy and yes, these methods are just as expensive as they sound. These methods often involve years of trial & error experimentation just to find even one protein structure. The number of protein structures currently determined by these methods is only a drop in the ocean when compared to all the possible protein targets for drugs that are out there.
You know what beats years? Days.
At the moment, the AlphaFold software takes only a few days to predict protein structure, and can predict the structure to the same degree of accuracy as all other methods – just the width of an atom’s worth of uncertainty. This allows scientists to predict the protein structure from the protein sequence with the same degree of accuracy as conventional methods, but in days not years.
AlphaFold's Predicted Structure (Blue)
vs
Experimentally Determined Structure (Green)

Image taken from the following source: AlphaFold: a solution to a 50-year-old grand challenge in biology | DeepMind
With AI powered protein structure determination at such a high level of accuracy, potential drug molecules could be found and tested on a much shorter timeframe.
With further development, this technology could revolutionise the drug design process
For now, this software is still in development, but work in the direction of reducing the needed computing power could make AlphaFold and softwares like it commonplace in labs and institutes working with such proteins.
For the scientists working in related fields, AlphaFold offers a glimmer of hope in somewhat troubling times.